Inflammation in severe asthma
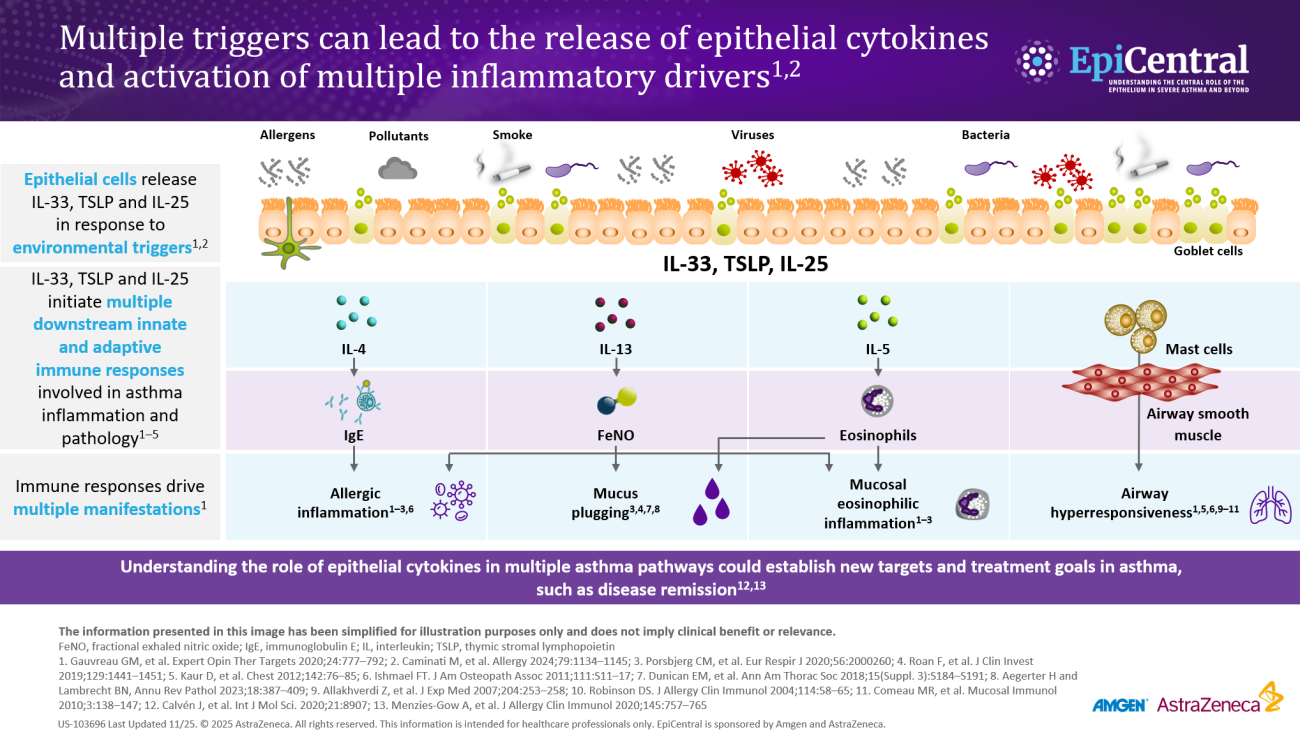
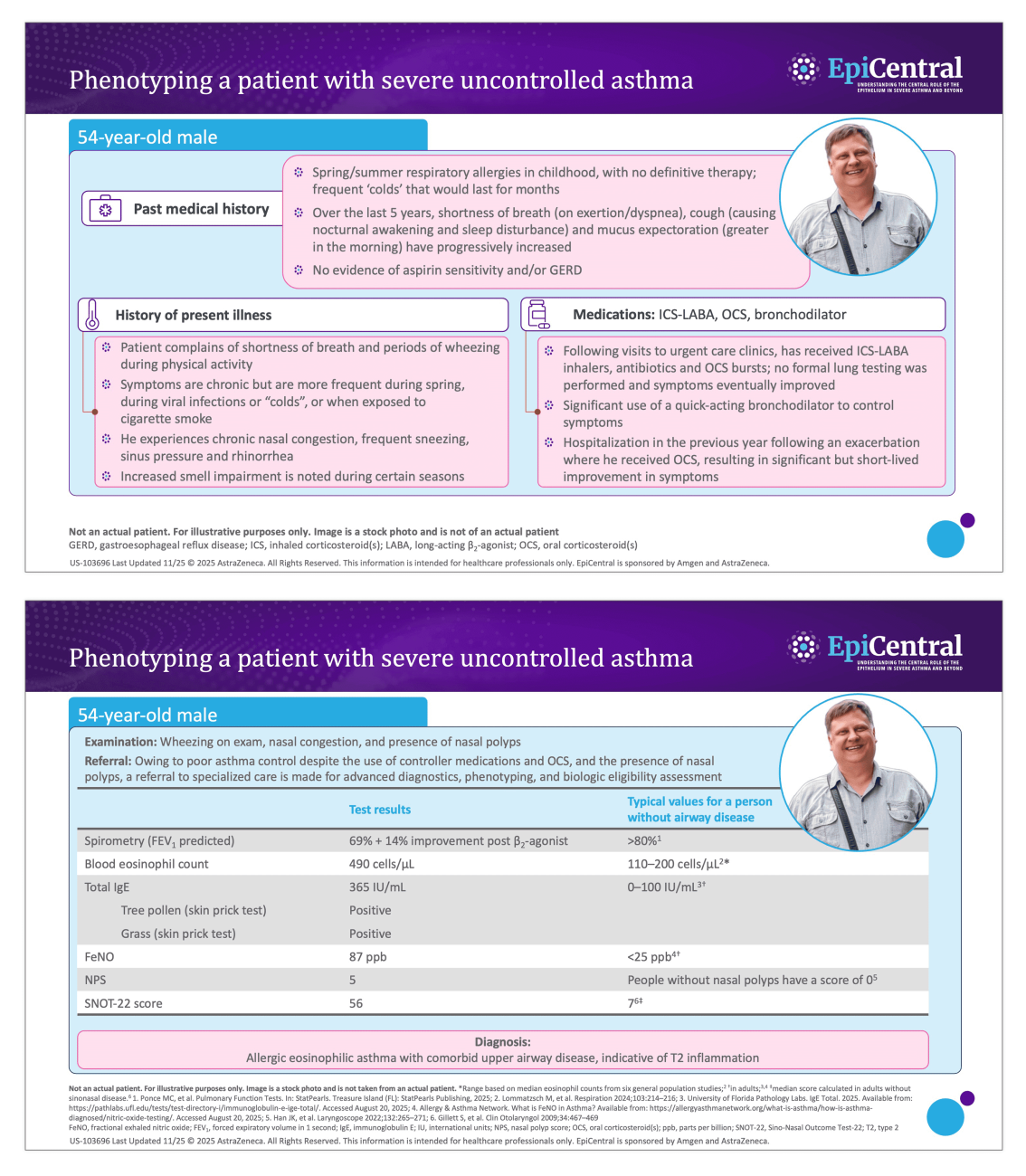
Asthma-associated inflammation is complex and heterogeneous,1–4 and numerous cell types, mediators, and downstream immune pathways are involved.1–6 Multiple inflammatory endotypes have been characterized, including allergic and eosinophilic inflammation.6